Search
Sharing Knowledge improves Knowledge... Knowledge should come at as less cost as possible.
Posts
Showing posts from July, 2015

Posted by
Varun C N
BtB#2- Viable but non-culturable
- Get link
- X
- Other Apps
Posted by
Varun C N
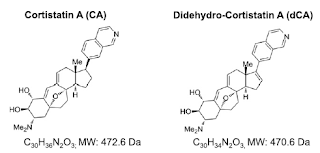
Attacking HIV Rebounce
- Get link
- X
- Other Apps
Posted by
Varun C N
BtB#1- Host parasite interaction interface
- Get link
- X
- Other Apps

Posted by
Varun C N
Engineered Microbes- They are the sensors
- Get link
- X
- Other Apps
Posted by
Varun C N
Break Annonucement
- Get link
- X
- Other Apps