Search
Sharing Knowledge improves Knowledge... Knowledge should come at as less cost as possible.
Posts
Showing posts from June, 2015

Posted by
Varun C N
Blogger's Desk #7: Guest post: Being a microbiologist: Roles and Responsibilities
- Get link
- X
- Other Apps
Posted by
Varun C N
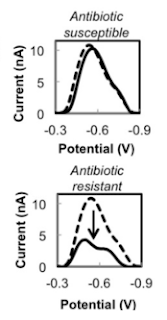
Chip makes a quick decision about antibiotics
- Get link
- X
- Other Apps

Posted by
Varun C N
MERS COV- Outbreak in Korea
- Get link
- X
- Other Apps

Posted by
Varun C N
Predatory Bacteria- Alternative to Antibiotics??
- Get link
- X
- Other Apps